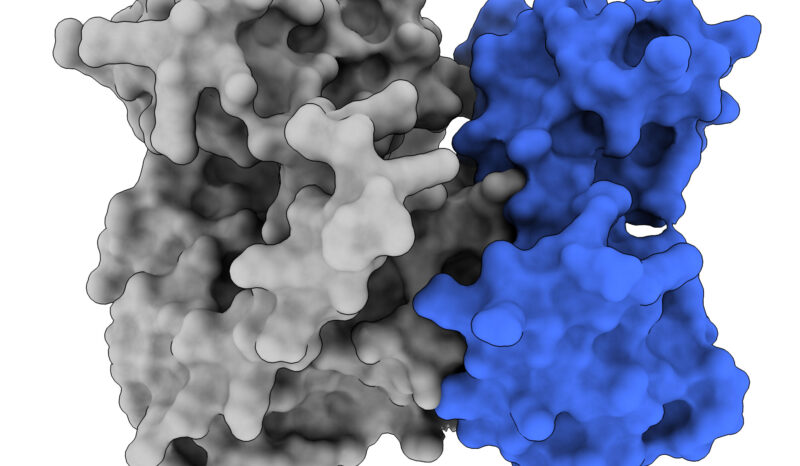
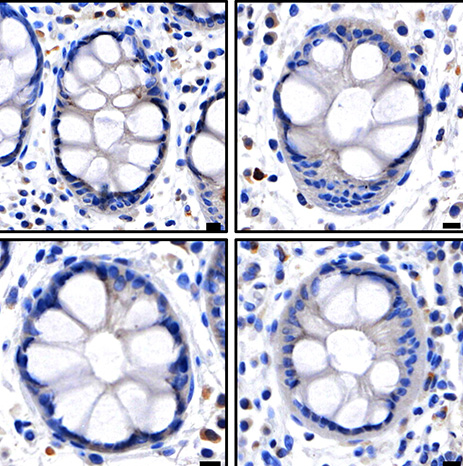
Our work has illuminated the molecular mechanism by which the killer protein, MLKL, is activated by the upstream kinase, RIPK3, to induce a pro-inflammatory form of cell death called necroptosis. Using innovative tools and integrative methods, we have defined four critical regulated steps in MLKL’s activation, which we term “checkpoints” in the necroptosis signaling pathway.
More broadly, we have validated pseudokinases, like MLKL, as underappreciated, but critical, signalling molecules throughout nature. Pseudokinases comprise ~10% of our kinome, and through detailed study, we have described diverse functional mechanisms by which pseudokinases function in signalling pathways, and how these functions are disrupted in disease. These studies have unearthed pseudokinases as novel candidates for therapeutic targeting.